The beginning of this new decade will mark a new open culture of research and collaboration, which are essential to make rapid advances in tackling COVID-19 (SARS-COV-2). This post looks at how aiLUNG computer simulations of coronavirus infection can drive research, including drug discovery.
Health Canada and the FDA perform a crucial function under normal circumstances by protecting the public from potentially dangerous drugs that have limited or no value to patients. The process of clinical development and approval is a robust, lengthy and costly undertaking for all involved. There are examples of very rapid conditional early approval of drugs based on compelling early clinical trial data. Gleevec for treating chronic myelogenous leukemia (CML) is only one such example, and there are many others, particularly when it comes to considering cancer therapies. The system can move very quickly when it is clear that it must.
The clinical drug development and approval process for new drug entities has three core elements
1. An approved treatment protocol based on earlier stage results.
2. A robust plan for patient safety and outcome assessment.
3. A detailed statement of potential risks and benefits to ensure informed consent.
Extraordinary times call for extraordinary (global) measures
Given the current global pandemic caused by COVID-19, I believe we are now living in extraordinary times, and we need to consider extraordinary measures when it comes to access to promising new drugs. In this regard, the issue of informed consent should emerge as the over-riding factor in determining access to potentially life-saving treatments. When a loved one is facing almost certain death should not the patient or family members be given the final decision about receiving a yet to be approved promising new treatment. This is where the principle of informed consent rather than approval should become the ultimate determinant of access. This approach would mean that licensing agencies around the world would have to work together to assess potential risk-benefit for the primary purpose of creating robust and informed consent documents explicitly written for the layperson. During extraordinary times like these, drugs approved in one jurisdiction from the developed world could be assessed from an informed consent perspective in all jurisdictions. Under normal circumstances, the siloed or ivory tower jurisdictional approach can make some sense but not during a global health catastrophe like the one that is upon us now.
Bottom Line – Global Informed Consent in a Global Pandemic
We live in extraordinarily difficult and uncertain times thanks to a global pandemic. Outside the box thinking involving a global response based on our comprehensive assessment of risk/benefit leading to robust informed consent may be one step towards the extraordinary response that is required. Perhaps our planning going forward could be to substitute the concept of global informed consent for the terms extraordinary or desperate measures.
Validating how computer simulations can speed up drug discovery for COVID-19
Over the last month, we’ve been working, almost exclusively, on developing COVID-19 simulations by first differentiating artificial pluripotent stem cells (aiPSC) into artificial lung cells (aiLUNG), and then infecting those with simulated COVID-19 virus. We then started looking at combinations of known drugs. We believe that drug combinations are a much more rational, logical and proven approach than single drugs for most serious diseases. This has important implications for COVID-19 by identifying novel and effective combinations of known drugs. The bigger long-range view of this would be that this technology should be developed so that it can deal with these outbreaks earlier in the future. When we experience another new virus and identify the genome, we can create simulations early on. With the validated genome, we can create simulations months ahead of the spread of the next lethal global pandemic. If six months ago, we had understood that this virus was going to be a significant health problem, and we probably did know, new technologies could have been brought to bear much earlier. In fact, the genome of SARS-COV-2 was published back in early 2020 and we had an early indication that the ACE2 receptor was the entry point to the cell. Looking at the near future, machine learning-based computer simulations can help to improve preparedness for future pandemics, which are certain to threaten us.
“Many people still don’t believe that it’s possible to create a simulation of a pluripotent stem cell in a computer. Well, we’ve done it. We’ve validated it. We have published and continuing to publish our results in peer-reviewed journals. Our validation is ongoing, and hopefully this and other technologies will, in time, be viewed as any other accepted lab test with positive and negative predictive values. When a new in-vitro test or animal model is developed, you don’t often get them right the first time.“
The first task is to find already approved drugs, whose clinical and side effect profiles are well known, that could be combined. Our current simulations are focused on inhibitors of RdRP, an RNA dependent RNA polymerase, and TMPRSS2, which is a cell surface receptor that cleaves part of the Spike protein to allow it to bind to the ACE2 receptor. When you combine hydroxychloroquine (HCQ) with either of those two inhibitors, you get quite dramatic and beneficial effects. If you next add an interleukin 6 inhibitor (IL-6), there are further statistically significant effects, particularly on the dangerous cytokine storm that occurs in about 30 percent of people who contract a COVID-19 infection.
Lastest aiLUNG results from COVID-19 simulations
New results from our ongoing COVID-19 project indicate that the combination of a TMPRSS2 inhibitor (Camostat mesylate) + hydroxychloroquine (HCQ) and an RdRP inhibitor (Remdesivir, Idarubicin) + HCQ should also be tested immediately.
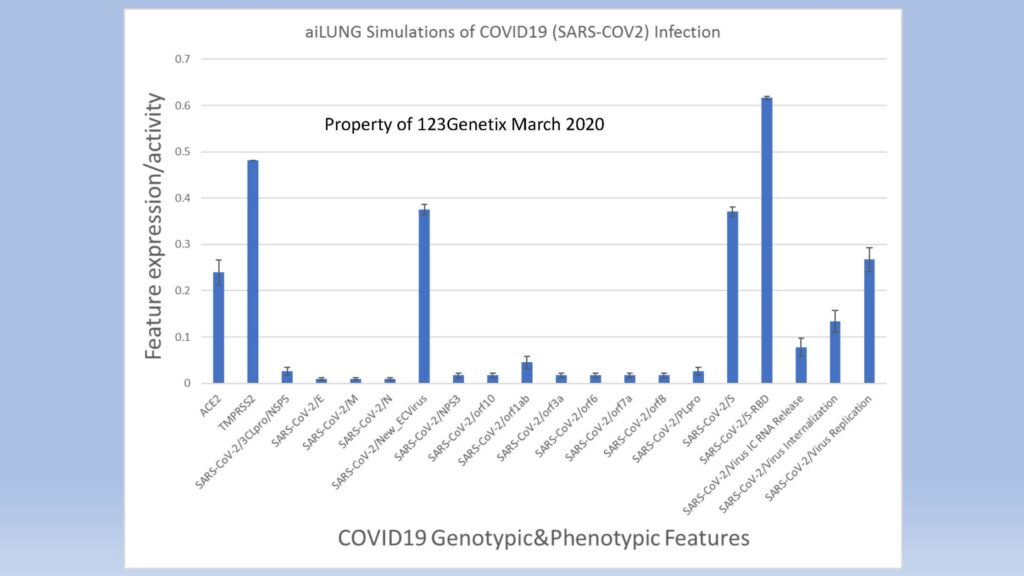
123Genetix has created and validated a simulated population of differentiated lung cells (aiLUNG) for the purposes of simulating COVID-19 infection. The validated COVID19 genome was simulated based on the most recent published data (March 2020). The aiLUNG cells with and without COVID-19 infection have been validated using the published literature. We are also now engaged in a preliminary screening and discovery program exploring single, double and triple-drug combinations of currently licenced drugs in Canada and the USA.
Our immediate goal is to optimize and finalize the aiLUNG-COVID-19 simulations based on domain expertise. We will then identify drug/drug combinations and take the potential candidates to in-vitro testing and animal models of the disease. Partners are required immediately to move this project towards effective therapies for the growing population of infected people around the globe.
It is our goal at this point to focus on identifying potential partners for continuing the research into and development of validated simulations of aiLUNG viral infections beginning with COVID-19/SARS-COV-2. The DeepNEU platform is flexible and can simulate many types of differentiated lung tissues that could be used to specifically further study the genetics of COVID-19.
The 123Genetix Rare Diseases Database v5.0 is freely available and aimed at rare diseases and now COVID-19 infection researchers and patient groups. You can see the sophistication of our modelling work here.
This is an unprecedented time when it is heartwarming to see research communities come together in a spirit of global innovation.
Featured image Fusion Medical Animation on Unsplash
Adapted from a LinkedIn article – Extraordinary times call for extraordinary (global) measures – March 27, 2020
Please give us your explicit consent to ensure that you continue to receive these blog notifications.